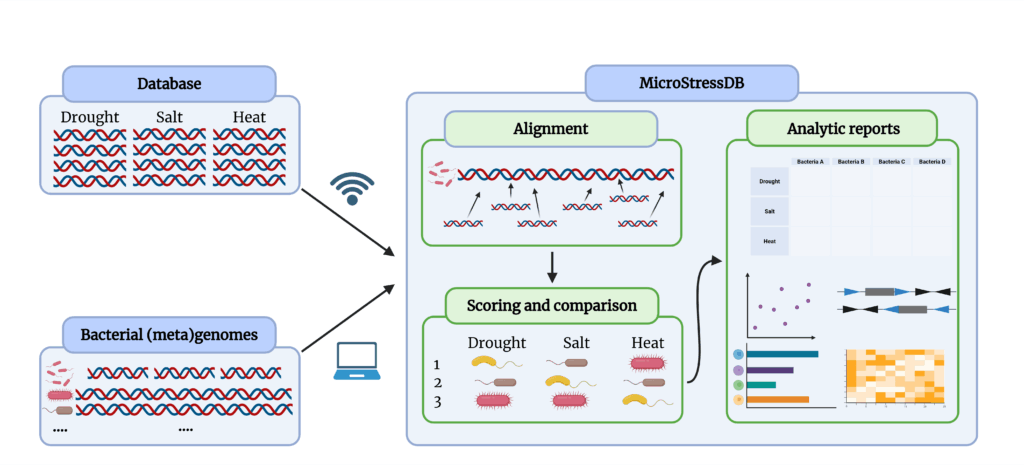
microSTRESS is an automated screening tool that scans the predicted proteome of a bacterium against a curated database of stress-alleviating gene families. It is designed for researchers working with plant-associated and environmental bacteria who want to rapidly identify genes involved in osmotic, oxidative, heat, and other stress-tolerance pathways.
Simply upload your protein FASTA file and enter your email address. The analysis runs on our server and you will receive a notification with a link to your personalised interactive HTML report — typically within a few minutes.
- Input: predicted proteome in protein FASTA format (
.faaor.fasta) — e.g. the.faaoutput from Prokka, RAST, or an NCBI genome annotation - Output: interactive HTML report with BLAST hit tables, identity & score charts, and a per-pathway summary
- Limit: max 100 MB per file
Your file is processed confidentially and automatically removed from our server within 24 hours. Only your email address and filename are retained for usage statistics.
💡 Know a stress gene that should be in MicroSTRESSdb? Suggest a new gene →
